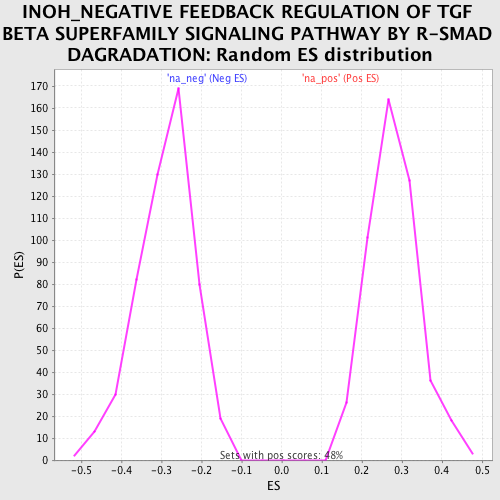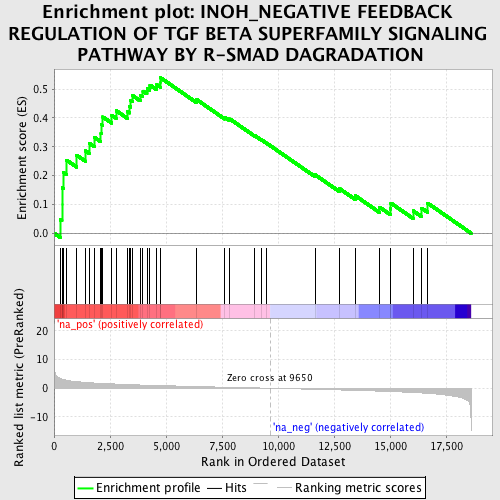
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | INOH_NEGATIVE FEEDBACK REGULATION OF TGF BETA SUPERFAMILY SIGNALING PATHWAY BY R-SMAD DAGRADATION |
| Enrichment Score (ES) | 0.54008776 |
| Normalized Enrichment Score (NES) | 1.9362657 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.043911785 |
| FWER p-Value | 0.289 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PSMB1 | 271 | 3.438 | 0.0475 | Yes | ||
| 2 | TLE4 | 353 | 3.194 | 0.1009 | Yes | ||
| 3 | TGFBR1 | 354 | 3.193 | 0.1585 | Yes | ||
| 4 | PSMB6 | 406 | 3.058 | 0.2111 | Yes | ||
| 5 | PSMB10 | 554 | 2.796 | 0.2537 | Yes | ||
| 6 | PSMB9 | 1005 | 2.291 | 0.2708 | Yes | ||
| 7 | SMAD5 | 1406 | 2.028 | 0.2859 | Yes | ||
| 8 | SMAD2 | 1568 | 1.934 | 0.3122 | Yes | ||
| 9 | PSMB2 | 1796 | 1.824 | 0.3329 | Yes | ||
| 10 | PSMB3 | 2090 | 1.688 | 0.3477 | Yes | ||
| 11 | PSMA5 | 2104 | 1.682 | 0.3774 | Yes | ||
| 12 | PSMB8 | 2176 | 1.657 | 0.4035 | Yes | ||
| 13 | PSMA3 | 2575 | 1.524 | 0.4096 | Yes | ||
| 14 | TLE3 | 2771 | 1.457 | 0.4254 | Yes | ||
| 15 | TLE6 | 3270 | 1.298 | 0.4221 | Yes | ||
| 16 | BMPR1A | 3362 | 1.271 | 0.4401 | Yes | ||
| 17 | SMAD3 | 3409 | 1.255 | 0.4603 | Yes | ||
| 18 | BMPR2 | 3490 | 1.229 | 0.4782 | Yes | ||
| 19 | PSMA4 | 3845 | 1.130 | 0.4796 | Yes | ||
| 20 | PSMB5 | 3966 | 1.100 | 0.4930 | Yes | ||
| 21 | ACVRL1 | 4147 | 1.059 | 0.5024 | Yes | ||
| 22 | PSMA6 | 4276 | 1.032 | 0.5142 | Yes | ||
| 23 | ACVR2A | 4585 | 0.962 | 0.5150 | Yes | ||
| 24 | TGFBR2 | 4739 | 0.928 | 0.5235 | Yes | ||
| 25 | PSMD2 | 4743 | 0.926 | 0.5401 | Yes | ||
| 26 | PSMA2 | 6367 | 0.596 | 0.4635 | No | ||
| 27 | SMAD7 | 7618 | 0.363 | 0.4028 | No | ||
| 28 | PSMB4 | 7842 | 0.322 | 0.3966 | No | ||
| 29 | SMAD4 | 8957 | 0.125 | 0.3389 | No | ||
| 30 | PSMA7 | 9241 | 0.077 | 0.3250 | No | ||
| 31 | PSMB7 | 9486 | 0.035 | 0.3125 | No | ||
| 32 | TLE2 | 11653 | -0.378 | 0.2028 | No | ||
| 33 | BMPR1B | 12733 | -0.587 | 0.1553 | No | ||
| 34 | SMAD1 | 13450 | -0.737 | 0.1300 | No | ||
| 35 | PSMA1 | 14534 | -1.013 | 0.0900 | No | ||
| 36 | PSMC1 | 15017 | -1.140 | 0.0847 | No | ||
| 37 | ACVR1 | 15033 | -1.143 | 0.1045 | No | ||
| 38 | SMAD9 | 16029 | -1.480 | 0.0777 | No | ||
| 39 | ACVR2B | 16407 | -1.637 | 0.0870 | No | ||
| 40 | TLE1 | 16668 | -1.763 | 0.1049 | No |